1) The core claim
Translation only works if the same codon reliably yields the same amino acid in practice. AARS enzymes make that reliability real: they attach (“charge”) the correct amino acid onto the correct tRNA.
A codon table doesn’t do anything by itself. The decisive step is charging: an AARS enzyme recognizes a specific tRNA and attaches the correct amino acid to its acceptor end, using ATP. That is where “meaning” is enforced.
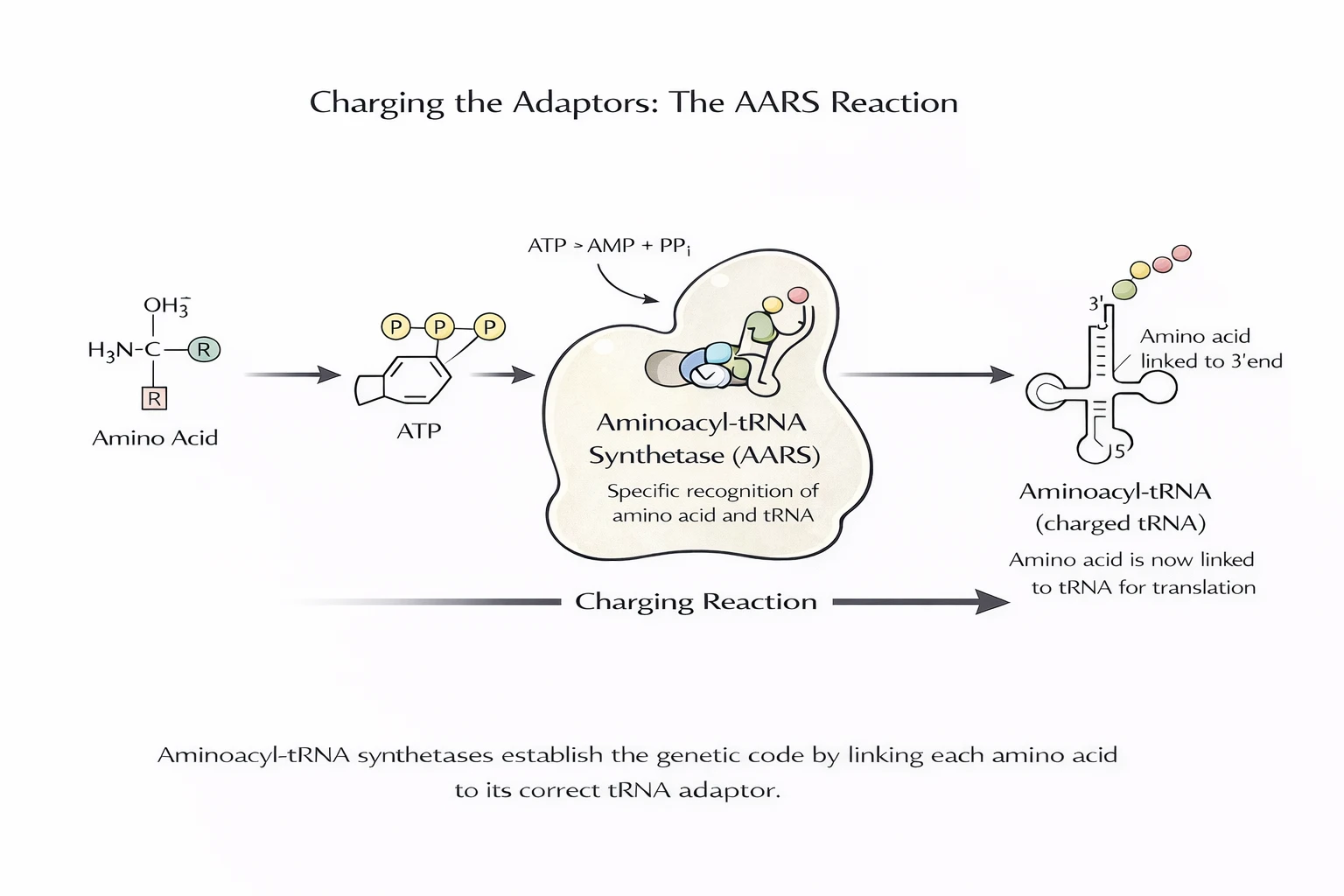
Charging alone is not sufficient. Many amino acids are chemically similar, so additional enforcement is required to prevent systematic mistranslation.
This is why many AARS enzymes include additional recognition and editing steps that actively reject incorrect substrates.

Think of the genetic code as a mapping that must be executed with low enough error to sustain replication and selection. The ribosome reads codons, but it does not “know” amino acids. It accepts whatever amino acid is carried by the tRNA that matches the codon. That means the genetic code is functionally implemented upstream at the charging step: AARS enzymes bind (i) an amino acid and (ii) the correct tRNA identity, then use ATP to covalently attach them.
If this enforcement fails—even modestly—translation becomes noisy, proteins misfold, and catalytic networks destabilize. From an origins standpoint, the key question becomes: how do you get a stable, self-maintaining mapping early enough for cumulative selection to operate on proteins at all?
The ribosome enforces reading. AARS enforce semantics (codon → amino acid) by establishing tRNA identity. Without charging, codons have no biochemical meaning.
Translation is a two-step constraint system: (1) the ribosome enforces a triplet reading frame; (2) AARS enforce the mapping between tRNA identity and amino acid identity.
tRNAs are not mere adapters; they are typed tokens. AARS enzymes are the type-checkers and (often) the proofreaders. They must distinguish among dozens of tRNAs, some of which share amino acids but use different anticodons.
In modern systems, fidelity emerges from layered constraints: identity elements in tRNA (acceptor stem, anticodon loop, discriminator base), AARS binding pockets tuned to amino acid chemistry, and—in certain cases—editing domains that hydrolyze near-cognate misactivations.
This is not just “an enzyme exists.” It is a distributed logic that must be coherent across: tRNA structures, AARS families, ATP-driven activation chemistry, ribosome acceptance rules, and downstream folding/quality control. The informational bottleneck is therefore not only a codebook, but the emergence of a stable enforcement network.