Translation System
How one-dimensional nucleotide sequences become three-dimensional catalytic structures — reliably, repeatably, and fast enough to outrun equilibrium.
Choose your level
This page is written in layers. Pick your preferred depth and reading style. (Nothing is tracked; this only changes what your browser shows.)
1) The core claim
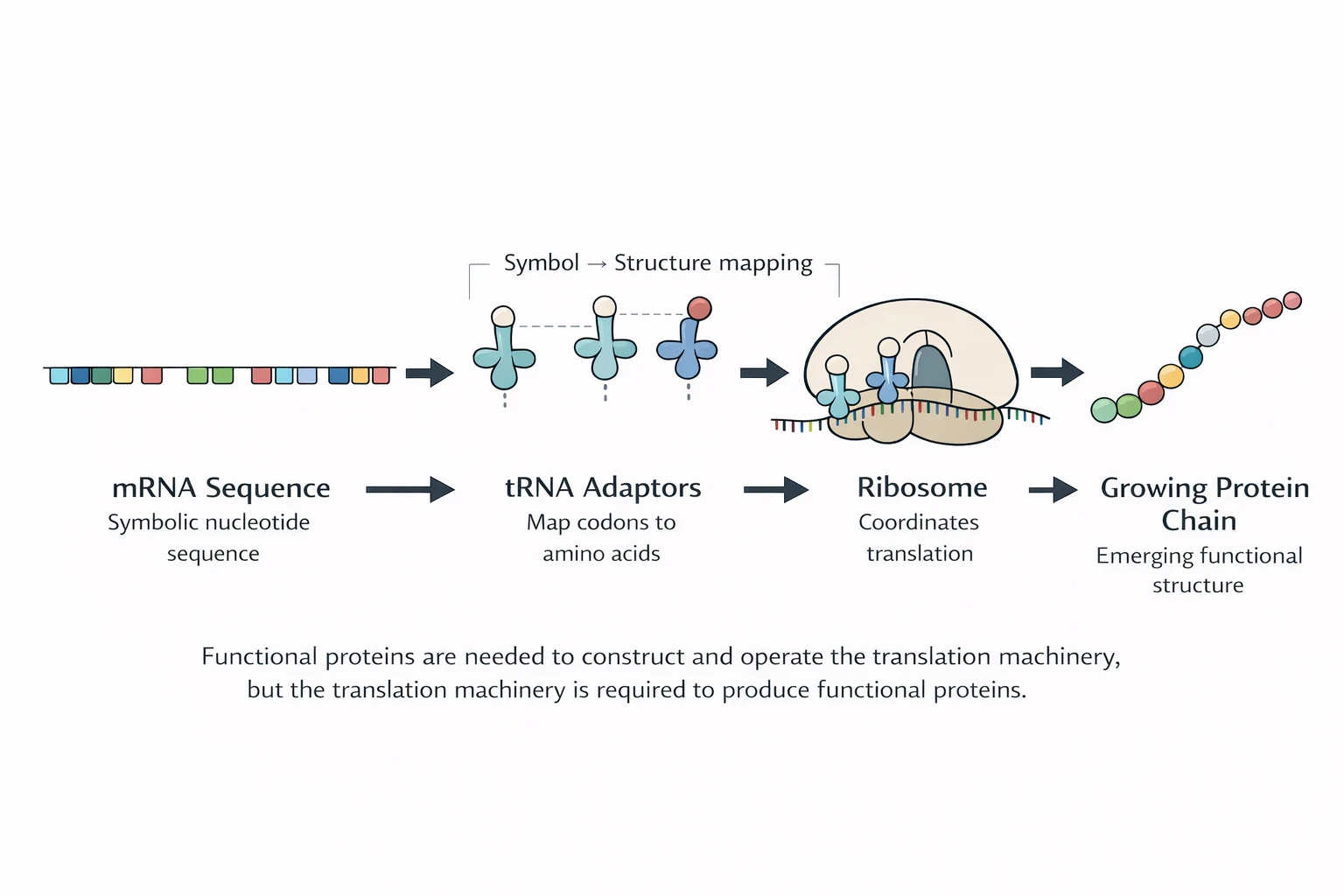
Translation is the step where “information” becomes chemistry: RNA code is read in triplets, and a protein chain is built in the matching order. Without translation, there is no enzyme production — and without enzymes, there is no stable metabolism or reproduction.
Translation is not merely “data processing.” It is a physical system that enforces a rule: codons (triplets) must be matched to amino acids consistently across the entire cell, under noise, drift, and damage.
This enforcement requires coordinated parts: ribosomes (reading frame control), tRNAs (adapters), aminoacyl-tRNA synthetases (AARS; charging + policing), initiation/elongation factors (timing + accuracy), and a downstream regime of folding, rescue, and degradation.
Translation is a coupled set of constraints: (1) maintaining a reading frame; (2) maintaining a stable codon→amino-acid mapping; (3) achieving throughput fast enough to sustain a non-equilibrium state; and (4) achieving fidelity high enough to avoid error catastrophe.
The system is therefore structurally and informationally closed: the machinery that enforces the mapping is itself a product of the mapping. This is the Phase-2 bottleneck that origins models must cross.
Working shorthand: genetic code = mapping, and translation = mapping + enforcement. Origins models can’t stop at “we have molecules that can store information.” They must explain how the mapping gets enforced quickly enough to persist.
2) The pipeline (what has to happen)
Step A — Charging: assign the correct amino acid to the correct tRNA
AARS enzymes attach amino acids to the right “carrier” RNAs (tRNAs). If the carrier is loaded wrong, the ribosome will still use it — and the protein will be wrong.
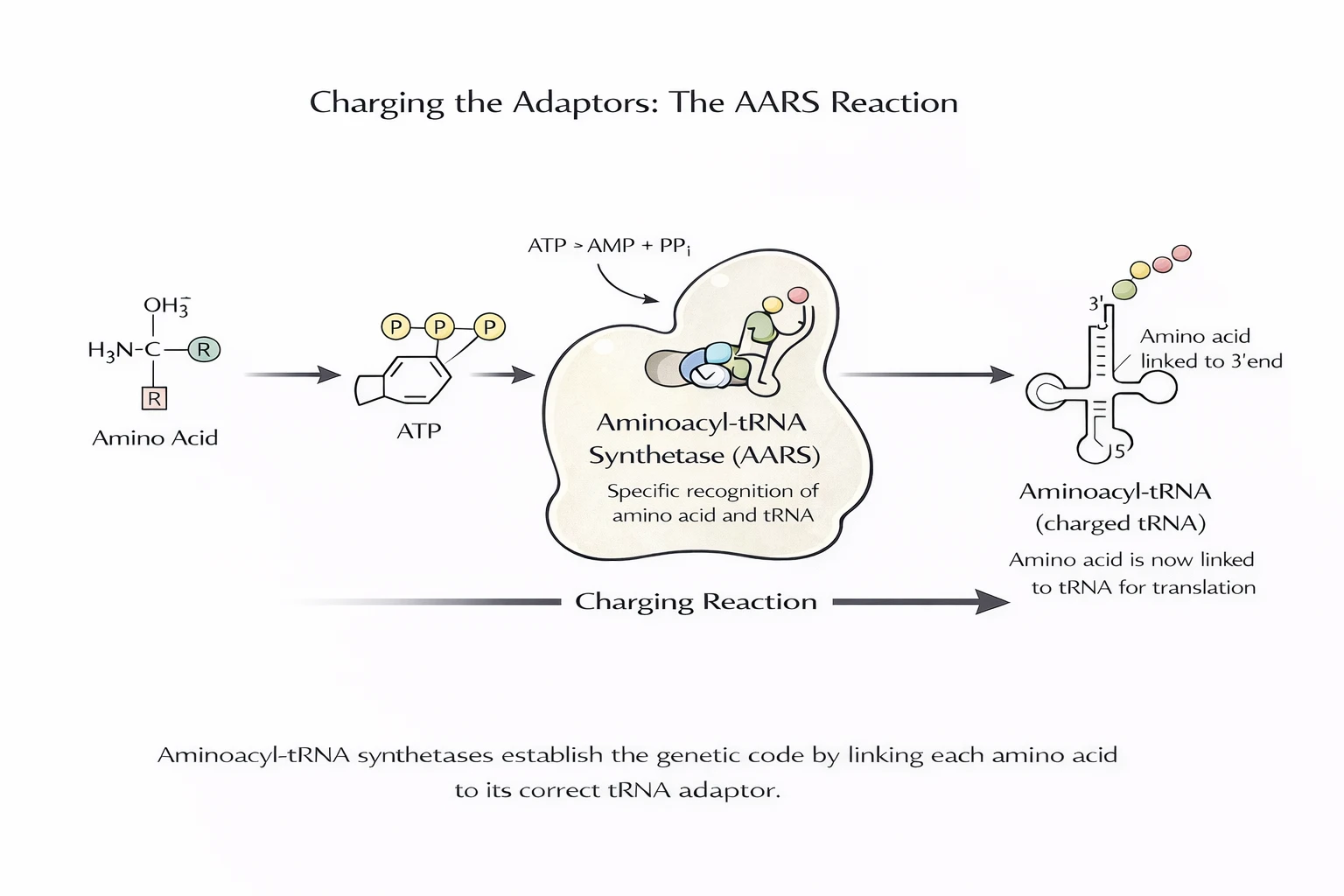
AARS enzymes “charge” tRNAs by attaching the appropriate amino acid to the tRNA’s acceptor end. This is where the mapping becomes real: the ribosome reads codons, but it does not “know” amino acids. It accepts whatever charged tRNA matches the codon.
Consequence: if a charged tRNA is wrong (misacylated), translation remains smooth — but wrong — producing proteins that can be nonfunctional, toxic, or destabilizing.
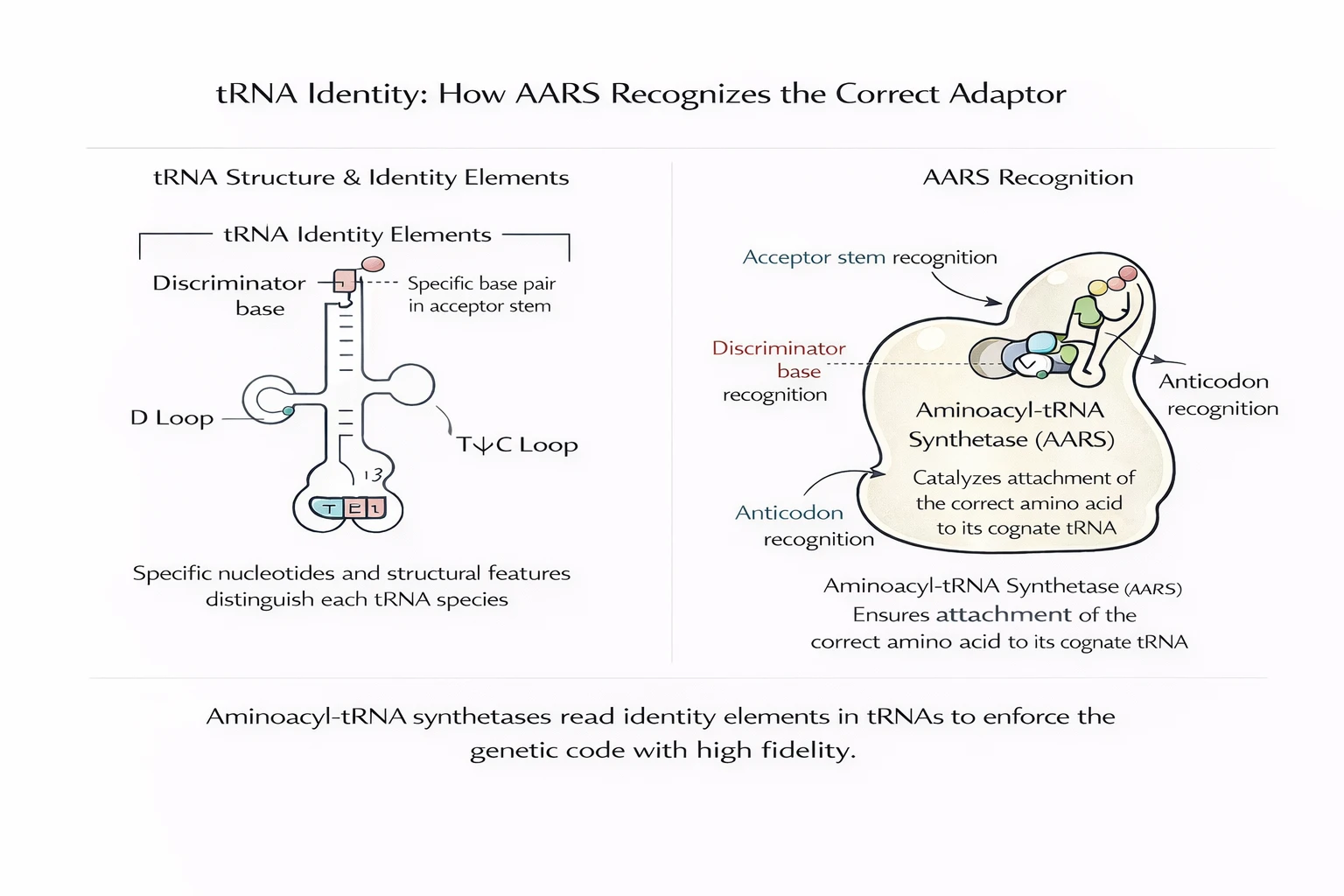
Mechanistically: AARS recognition uses structural features of tRNA (identity elements), sometimes including anticodon bases, discriminator bases, and acceptor stem geometry. Several AARSs have editing/proofreading functions to correct near-cognate amino acids.
This is why the AARS family is a “code enforcer” class: they instantiate the mapping in chemistry by repeatedly performing charged assignments with low error rates — at scale.
Step B — Initiation: start at the right place, in the right frame
The ribosome has to start reading at the right “start point.” If it starts one letter off, everything downstream changes.
Initiation factors help assemble the initiation complex: ribosomal subunits, mRNA, and the initiating tRNA align so the reading frame is defined. A one-nucleotide offset is catastrophic because every codon downstream is changed.
Reading frame integrity is an algorithm implemented in structure: ribosomal geometry and kinetic checkpoints reduce frameshifting. Origins models must explain how a stable frame-enforcing architecture can arise and persist.
Step C — Elongation: add amino acids quickly and accurately
For each codon, the ribosome selects a matching tRNA and adds its amino acid to the growing chain.
Elongation is enforced by the ribosome’s internal geometry. Incoming tRNAs are processed through three functional stations: the A site (aminoacyl-tRNA entry), P site (peptidyl-tRNA holding the growing chain), and E site (exit of the deacylated tRNA).
Peptide bond formation occurs in the ribosome’s rRNA catalytic core, followed by translocation: the ribosome advances exactly one codon, shifting tRNAs A → P → E in a tightly controlled cycle.
Elongation factors and ribosomal checkpoints bias toward correct codon-anticodon pairing. Speed matters because the cell must produce enough proteins fast enough to sustain itself, but speed competes with accuracy.
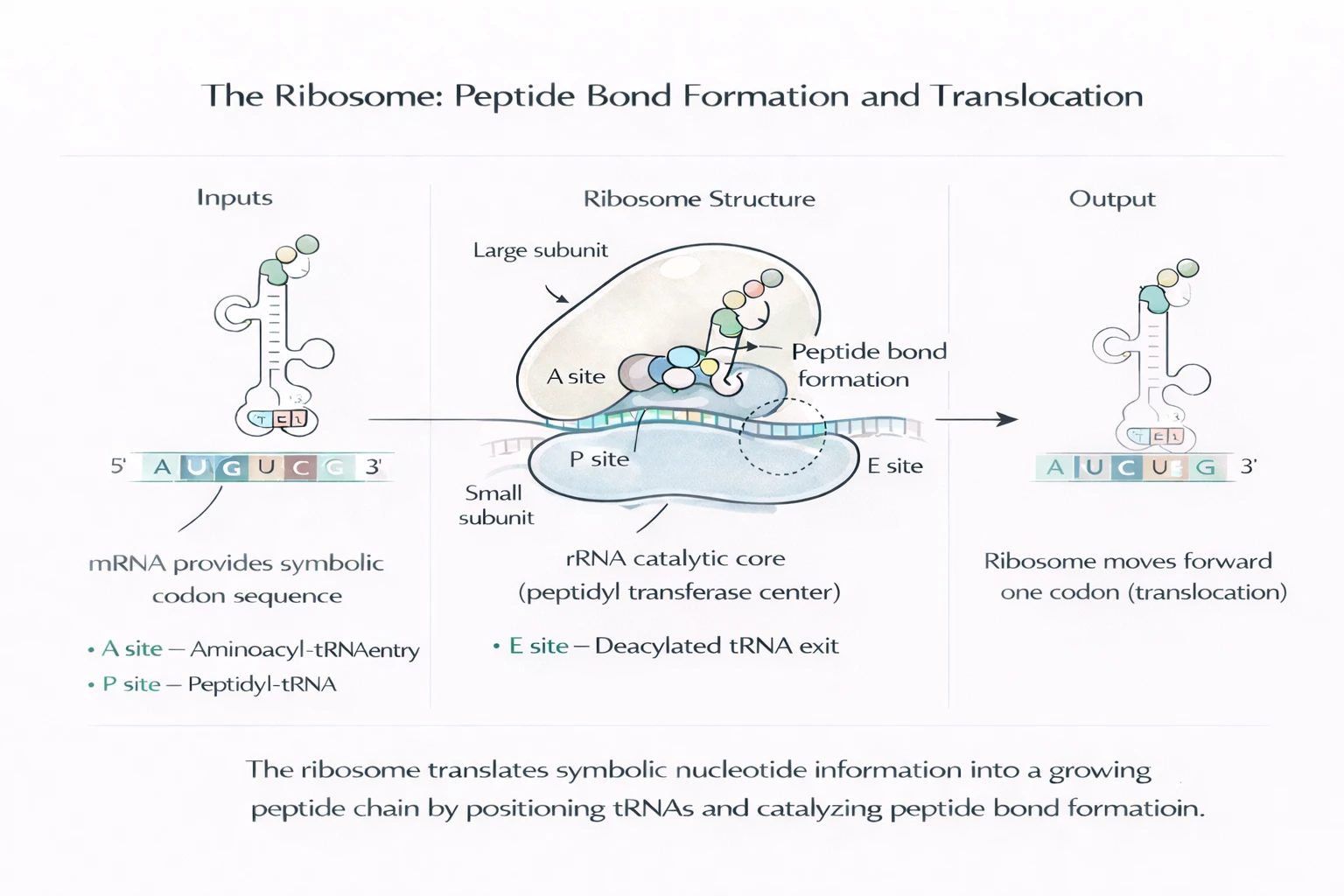
Detailed mechanics: Explore the ribosome →
Translation is therefore a throughput/fidelity trade space, not a simple “look-up table.”
Elongation is governed by kinetic proofreading, tRNA selection dynamics, and ribosome translocation. Errors include near-cognate acceptance, frameshifts, and premature termination.
Step D — Termination + release: stop at the right time
Stop codons signal the end. Release factors free the completed chain.
Termination depends on recognizing stop codons and releasing the polypeptide cleanly. Premature termination produces truncated proteins; failure to terminate causes jams and waste.
Termination and ribosome recycling are part of the system’s resource economy: stalled ribosomes must be rescued, components reused, and errors contained.
Step E — Folding, rescue, and quality control
New protein chains must fold into useful shapes. Many need help.
Folding is not automatic in crowded cytosol. Chaperones, refolding systems, and degradation pathways prevent aggregation and remove failures.
This “downstream policing” means translation is embedded in a broader control regime — a point that matters when thinking about how any early system could survive.
Quality control spans cotranslational folding, chaperone networks, proteolysis, and post-translational modifications. The minimal viable translation system is therefore larger than the ribosome alone.
3) Why this is a bottleneck for origins models
Any origins story has to explain how translation could appear early enough to matter. Without translation, you can’t make enzymes. Without enzymes, you can’t stabilize the chemistry.

Origins models can’t stop at “RNA can store information” or “ribozymes can catalyze something.” The central requirement is a system that reliably maps code to structure and can reproduce the mapping apparatus before drift and damage erase it.
That’s why Phase 2 is not optional glue between “origins” and “evolution.” It is the bridge that determines whether cumulative selection can ever get started.

The core dilemma is circular dependence: translation machinery is encoded by code that requires translation machinery to be expressed, maintained, and selected. “Partial systems” face discontinuities: missing one assignment can halt synthesis, while misassignments can poison the proteome.