3) Sparse function landscapes (why “most sequences” are noise)
Proteins only work when they fold into stable shapes. Most random amino-acid sequences don’t fold into anything useful—so “function” sits on rare islands inside a vast sea of noise.
Enzymes work because a sequence produces a specific 3D fold—a tightly constrained pattern of attractions and repulsions (hydrophobic packing, charge interactions, hydrogen bonds) that yields a stable structure and, sometimes, an active site.
This creates a harsh asymmetry: changing the sequence usually changes the fold, and a broken fold usually means lost function. In other words, the search space is dominated by sequences that are unstable, misfolded, aggregated, or non-catalytic. Functional proteins occupy sparse “islands” in sequence space rather than a smooth landscape.
Why this matters for bootstrapping
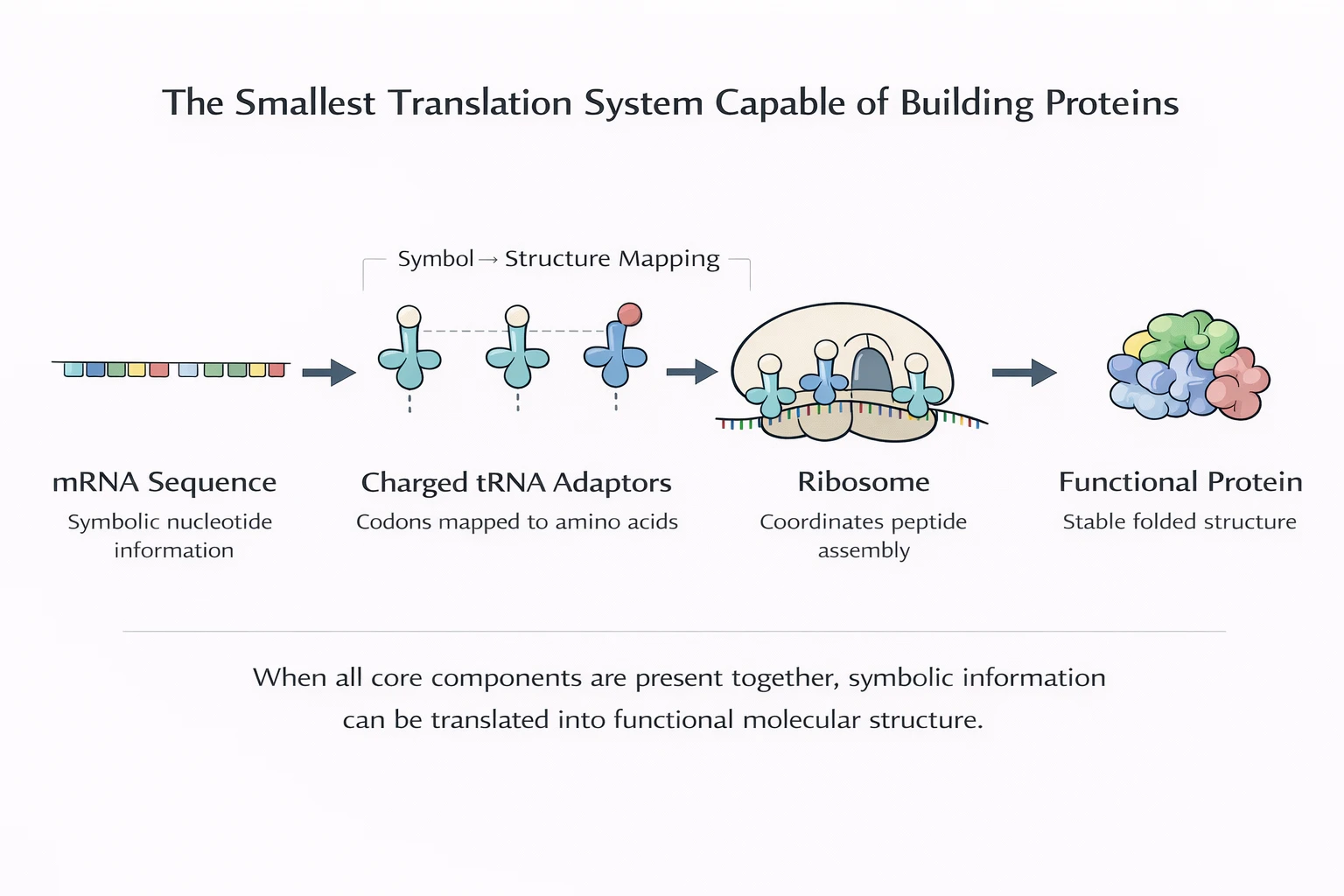
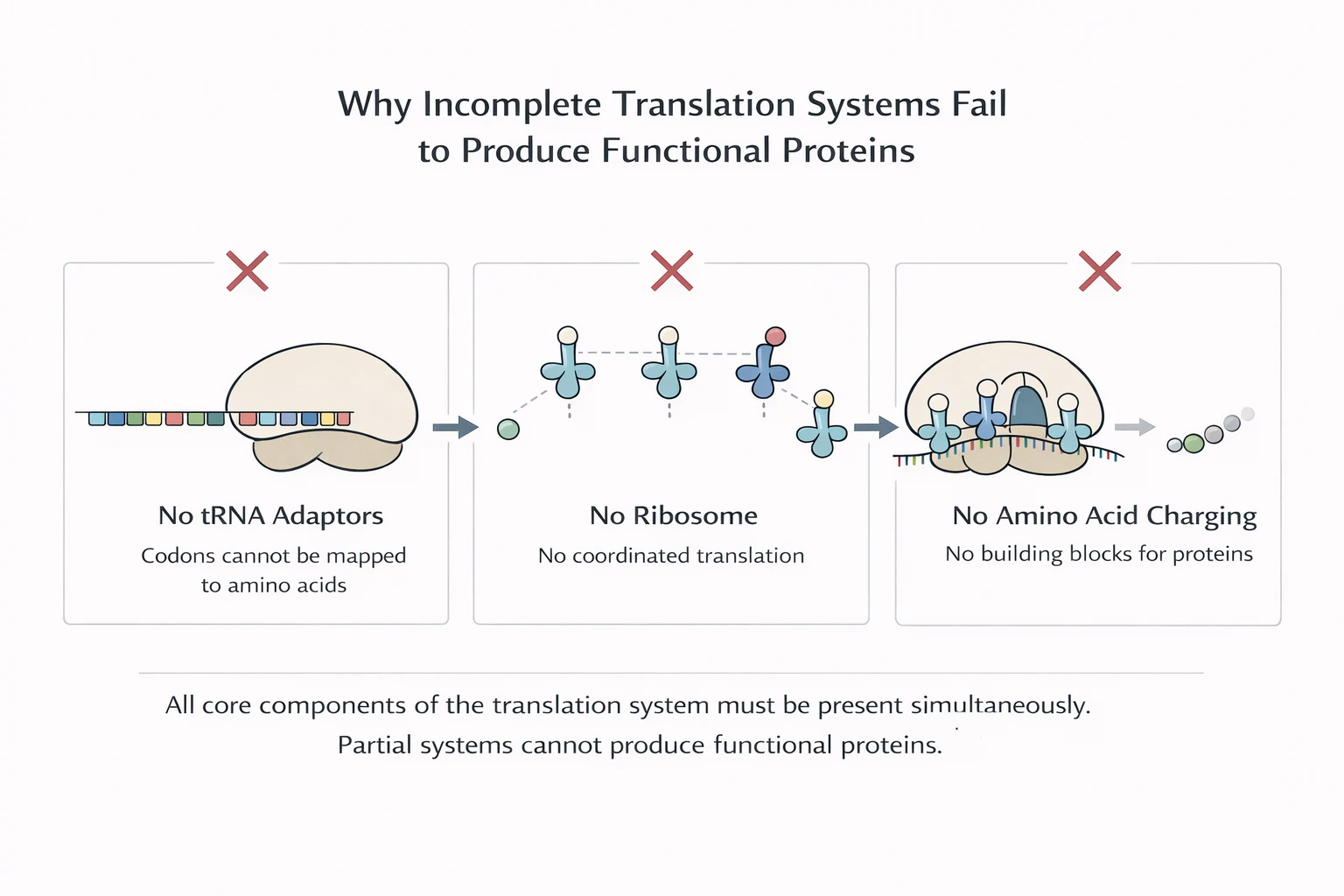
A translation system doesn’t just need to exist—it must reliably produce proteins that land in the functional regime often enough to sustain and rebuild the system itself.
The constraint is not merely combinatorial (“many possible sequences”), but structural: function is mediated through fold stability and geometry. Most sequences fail to form a cooperative, kinetically accessible fold, or they form folds that lack catalytic features. Small substitutions can rewire interaction networks, destabilize cores, or distort active-site geometry—often collapsing activity rather than smoothly degrading it.
This makes early evolution especially demanding. Before a lineage has reliable translation and repair, the system must repeatedly sample sequences—but the probability mass is concentrated in nonfunctional space. A viable pathway therefore needs both (a) a way to generate variation and (b) a way to retain rare gains long enough for cumulative improvement (i.e., closure under replacement and turnover).
Working takeaway
Phase 2 is a double threshold: closure (self-maintaining translation) inside a sparse functional landscape (rare stable catalytic folds).